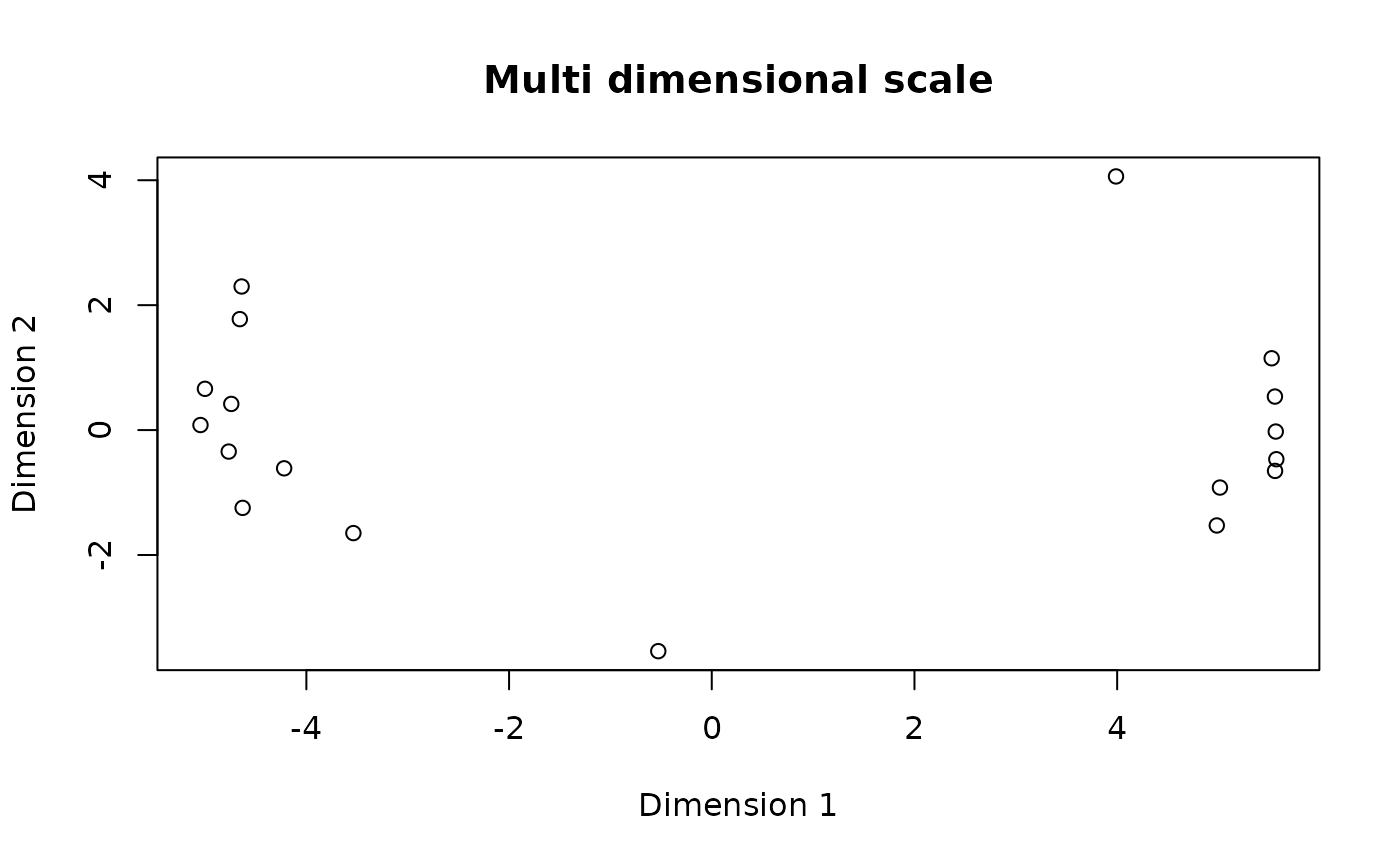
Convert salient sequences into MDS space
salient_mds(.mp, data, bit_idx = 1)Arguments
- .mp
a Matrix Profile object.
- data
the data used to build the Matrix Profile, if not embedded.
- bit_idx
an
int. The index ofn_bitsused for MDL discretization if more than one was used. (Default is1).
Value
Returns X,Y values for plotting
References
Yeh CCM, Van Herle H, Keogh E. Matrix profile III: The matrix profile allows visualization of salient subsequences in massive time series. Proc - IEEE Int Conf Data Mining, ICDM. 2017;579-88.
Hu B, Rakthanmanon T, Hao Y, Evans S, Lonardi S, Keogh E. Discovering the Intrinsic Cardinality and Dimensionality of Time Series Using MDL. In: 2011 IEEE 11th International Conference on Data Mining. IEEE; 2011. p. 1086-91.
Examples
# toy example
data <- mp_toy_data$data[, 1]
mp <- tsmp(data, window_size = 30, verbose = 0)
mps <- salient_subsequences(mp, verbose = 0)
mds_data <- salient_mds(mps)
plot(mds_data, main = "Multi dimensional scale")